EPJ E Highlight - Considering how friction is maximised when liquids flow on nanoscales
- Details
- Published on 15 August 2022

By simulating a liquid confined by a nanoscale structure, researchers discovered the role molecular clogging plays in friction.
The dynamics of how liquids behave when confined in a nanoscale-sized space such as nanochannels, nanotubes or nanopores, is key to understanding a wealth of processes including lubrication, filtration and even energy storage.
The dynamics of liquids at nanoscales are different to behaviour in confinement at macroscales, however. One of the key differences that a reduction in scale creates is friction and shear between the liquid and its solid container. And further complications arise in systems with solid-to-solid contact with features like wear, micro-pitting and scuffing created.
A new paper published in EPJ E and authored by Shan Chen, from the State Key Laboratory of Organic-Inorganic Composites at Beijing University of Chemical Technology, China, uses simulations of molecular dynamics to look at the friction-induced nano-confined liquids.
EPJ E Colloquium - Predicting thermodiffusion in simple binary fluid mixtures
- Details
- Published on 14 June 2022

When a homogeneous mixture is subjected to a thermal gradient, the fluid components are partially separated because of the temperature gradient. This phenomenon, known since the mid-19th century, is called thermodiffusion, the Soret effect or thermophoresis. Despite its relatively small amplitude it impacts many natural systems, such as the salinity gradient in ocean or even pre-biological evolution, and can be exploited in applications ranging from the manipulation of biological macromolecules to isotope enrichment. However, despite numerous attempts by leading researchers, including some Nobel laureates, a full understanding of the microscopic origin of this subtle phenomenon is still lacking and there is no consensus on which model, among the numerous existing ones, is the most reliable to quantify it in dense phases.
EPJ E Highlight - Modelling the behaviour and dynamics of microswimmers
- Details
- Published on 24 May 2022

The understanding of the clustering and movement of microswimmers has a range of applications from human health to tackling ecological problems.
Microswimmers are biological entities that range from sperm to phytoplankton to bacteria, meaning that their study can have implications for fields in science as diverse as human health and ecology.
A new paper published in EPJ E looks at the dynamics of microswimmers under gravity. It is authored by a team from the Institute for Theoretical Physics at the Berlin Institute of Technology: Felix Rühle, Arne W. Zantop, and Holger Stark.
EPJ E Highlight - The relationship between active areas and boundaries with energy input in snapping shells
- Details
- Published on 05 April 2022

New research looks at how the geometry of shells relates to the energy input required to actuate snap-through instability.
In nature, diverse organisms such as the hummingbird and Venus flytrap use rapid snapping motions to capture prey, inspiring engineers to create designs that function using snap-through instability of shell structures. Snapping rapidly releases stored elastic energy and does not require a continuously applied stimulus to maintain an inverted shape in bistable structures.
A new paper published in EPJ E authored by Lucia Stein-Montalvo, Department of Civil and Environmental Engineering, Princeton University, and Douglas P. Holmes, Department of Mechanical Engineering, Boston University, along with co-authors Jeong-Ho Lee, Yi Yang, Melanie Landesberg, and Harold S. Park, examines how restricting the active area of the shell boundary allows for a large reduction in its size, and decreases the energy input required to actuate snap-through behaviour in the shell to guide the design of efficient snapping structures.
EPJE has appointed new Editor-in-Chief Giovanna Fragneto
- Details
- Published on 08 December 2021

The publishers of European Physical Journal E: Soft Matter and Biological Physics are delighted to announce the appointment of Prof Giovanna Fragneto as Editor-in-Chief, starting January 1 2022. Prof Fragneto has served on the Editorial Board of EPJE since 2011, and takes over the EiC role from Prof François Graner, who steps down at the end of this year.
Prof Fragneto joins Prof Fabrizio Croccolo and Prof Holger Stark as Editors-in-Chief for EPJE, with collective responsibility for papers submitted across the scope of the journal.
EPJE Topical review - Advances in the study of supercooled water
- Details
- Published on 29 November 2021
EPJ E Topical review - Structure and dynamics of nanoconfined water and aqueous solutions
- Details
- Published on 16 November 2021
Water, regarded as the matrix of life, is an ubiquitous and peculiar liquid that exhibits a plethora of anomalous properties, both in its stable and metastable bulk states, which fostered a lot of experimental and theoretical studies. Less explored is the field of water and aqueous systems confined in nanoporous materials that, in addition to its fundamental interest, are present in a number of practical situations, including biological and separation processes and energy generation and storage, among others. These facts have triggered a vast amount of research that, so far, has not been conveniently reviewed.
EPJ E Highlight - Simulating microswimmers in nematic fluids
- Details
- Published on 12 July 2021
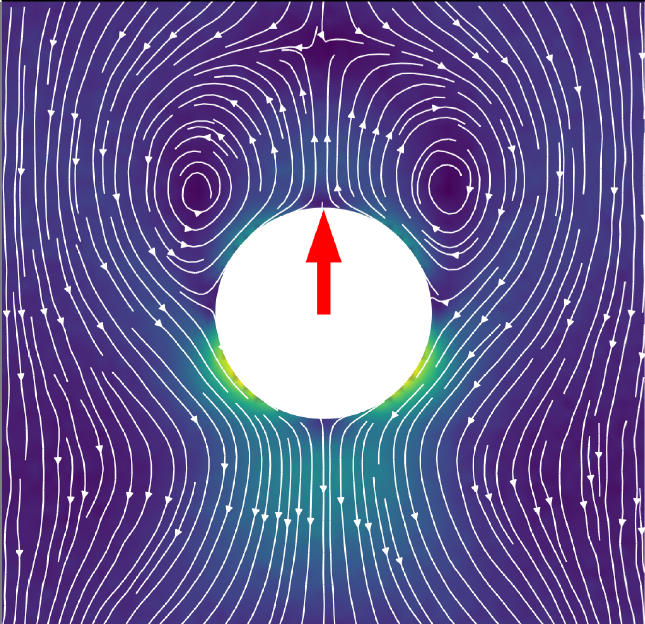

A combination of two simulation techniques has allowed researchers to investigate how swimming microparticles propel themselves through ‘nematic liquid crystals’ – revealing some unusual behaviours
Artificial microswimmers have received much attention in recent years. By mimicking microbes which convert their surrounding energy into swimming motions, these particles could soon be exploited for many important applications. Yet before this can happen, researchers must develop methods to better control the trajectories of individual microswimmers in complex environments. In a new study published in EPJ E, Shubhadeep Mandal at the Indian Institute of Technology Guwahati (India), and Marco Mazza at the Max Planck Institute for Dynamics and Self-Organisation in Göttingen (Germany) and Loughborough University (UK), show how this control could be achieved using exotic materials named ‘nematic liquid crystals’ (LCs) – whose viscosity and elasticity can vary depending on the direction of an applied force.
EPJ E Highlight - Micro-environmental influences on artificial micromotors
- Details
- Published on 31 March 2021
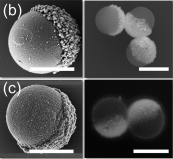

New experiments reveal the characteristic ways in which self-propelled ‘Janus particles’ with charged coatings will slide across or move away from charged boundaries in their surrounding environments.
By harvesting energy from their surrounding environments, particles named ‘artificial micromotors’ can propel themselves in specific directions when placed in aqueous solutions. In current research, a popular choice of micromotor is the spherical ‘Janus particle’ – featuring two distinct sides with different physical properties. Until now, however, few studies have explored how these particles interact with other objects in their surrounding microenvironments. In an experiment detailed in EPJ E, researchers in Germany and The Netherlands, led by Larysa Baraban at Helmholtz-Zentrum Dresden-Rossendorf, show for the first time how the velocities of Janus particles relate to the physical properties of nearby barriers.
EPJ E Highlight - Modelling speed-ups in nutrient-seeking bacteria
- Details
- Published on 17 March 2021
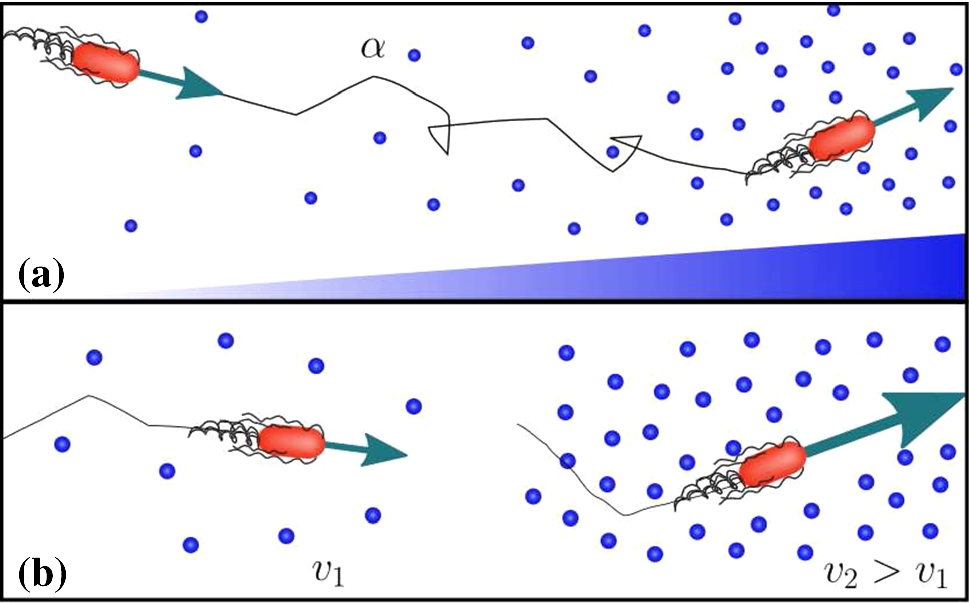
By considering how some bacteria will swim faster within higher nutrient concentrations, researchers have created a more accurate model of how these microbes search for nutrients
Many bacteria swim towards nutrients by rotating the helix-shaped flagella attached to their bodies. As they move, the cells can either ‘run’ in a straight line, or ‘tumble’ by varying the rotational directions of their flagella, causing their paths to randomly change course. Through a process named ‘chemotaxis,’ bacteria can decrease their rate of tumbling at higher concentrations of nutrients, while maintaining their swimming speeds. In more hospitable environments like the gut, this helps them to seek out nutrients more easily. However, in more nutrient-sparse environments, some species of bacteria will also perform ‘chemokinesis’: increasing their swim speeds as nutrient concentrations increase, without changing their tumbling rates. Through new research published in EPJ E, Theresa Jakuszeit and a team at the University of Cambridge led by Ottavio Croze produced a model which accurately accounts for the combined influences of these two motions.




